
1. Introduction: Why Genes Need Regulation

Escherichia coli (E. coli, for short) is an intestinal bacterium we’ve met several times in this course. As you’ll remember, it inhabits the human colon, or large intestine, as well as the colons of many other mammals.
E. coli’s genome consists of over 4000 protein-coding genes(Wikipedia). While some of these proteins are constantly in use, others are only useful under certain conditions. For example, E. coli possesses a group of enzymes that it uses to digest the disaccharide lactose (the sugar in milk). If E. coli is in a lactose-rich environment, expressing the genes for lactose-digesting enzymes will enable it to convert lactose into monosaccharides that can be further broken down during the reactions of cellular respiration. But if there’s no lactose, then producing the enzymes for lactose digestion would be a waste of energy. So, it’s of adaptive value for E. coli to be able to control its production of these enzymes to match the availability of lactose in the environment. How does E. coli do this?
2. Operons: Description and Definition
Many of E. coli’s genes (as well as many genes in other bacteria, archaea, viruses, and, more rarely, eukaryotic organisms) are organized into systems called operons. Operons consist of the following components.
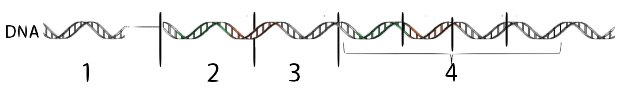
The DNA at “4” consists of structural genes. These are the genes that code for enzymes or other gene products. These structural genes are under the control of a regulatory gene (1). The regulatory gene produces a regulatory protein that determines whether or not the structural genes will be transcribed into mRNA, which in turn is translated into protein. Because the regulatory protein often has the effect of preventing transcription, the regulatory protein is often referred to as a repressor.
Transcription is carried out by RNA polymerase, which can only transcribe a gene by binding with a region of DNA called the promoter, shown above at “2.” Just downstream of the promoter is a region called the operator (3). The operator is a binding site for the regulatory protein/repressor that the regulatory gene codes for. Pulling this together, we can define an operon as follows (modified from Wikipedia):
…a cluster of genes under the control of a single promoter. The genes are transcribed and regulated in such a way that these genes are either expressed together or not at all.
This will become clearer by looking at an operon in action. We’ll start with the lac operon, the first operon whose function was understood. This understanding came about through the work of François Jacob, André Michel Lwoff, and Jacques Monod, who received the Nobel Prize for their work in 1965.
3. The Lac Operon
The Lac operon is the cluster of structural genes described above: they code for a series of enzymes that work together to convert lactose into two monosaccharides: glucose and galactose. Here’s how the expression of these structural genes is controlled.
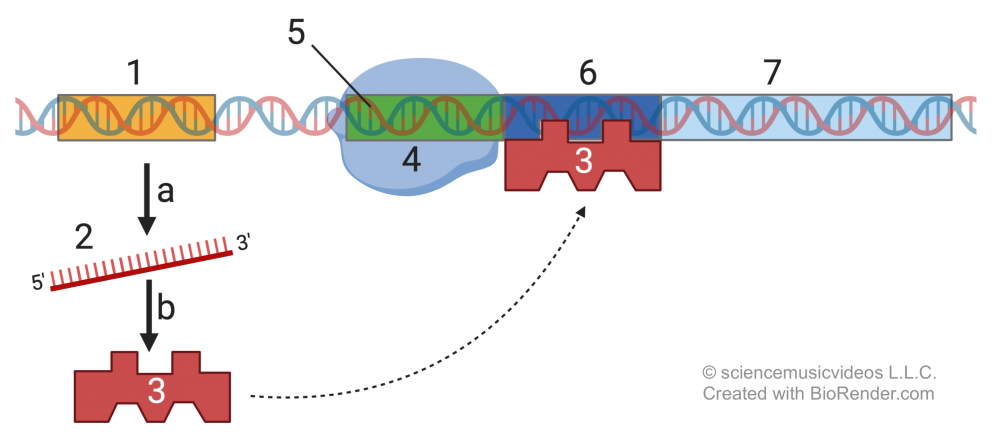
The DNA at “1” is the regulatory gene. This gene is transcribed (process “a”) into mRNA (2), which in turn is translated (process “b”) into the regulatory protein (3).
The regulatory protein (also known as a repressor) has two binding sites. In the absence of lactose, one of its binding sites is complementary to the operator region (6) of the Lac operon. When the regulatory protein binds to the operator, RNA polymerase (4) can’t bind with its promoter (5), and, as a result, the structural genes (7) can’t be transcribed.
Below, you can see how the situation changes when lactose is in the environment.
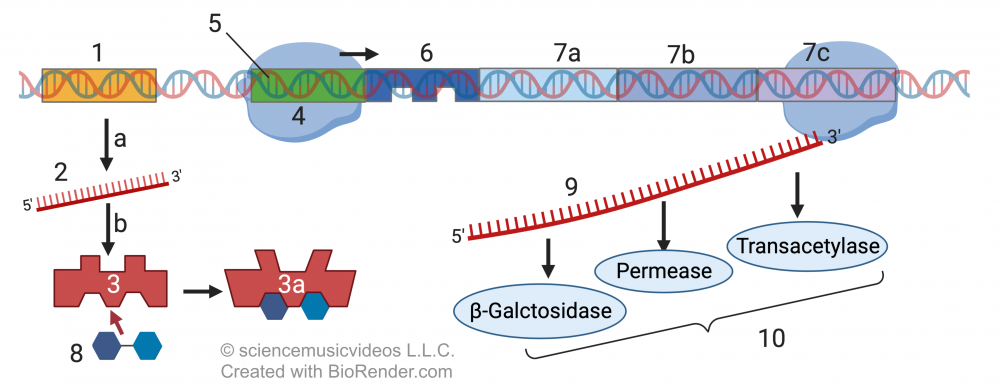
Lactose (8) will bind with the regulatory protein in a way that causes an allosteric shift so that the regulatory protein’s DNA binding site is no longer complementary to the lac operon’s operator (6). As a result, RNA polymerase can bind with its promoter and can transcribe the structural genes (7a, 7b, and 7c) into mRNA (9), which can, in turn, be translated into the enzymatic proteins that digest lactose.
4. Lac Operon: Labeled Diagrams
Got it? Prove it by labeling the interactive diagrams below.
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Lac Operon Labeled Diagrams (M16)”]
[h]Lac Operon: Interactive Diagrams
[i] Biohaiku
Operon System
Controls expression of genes
Such efficiency!
[q labels = “top”]
[l]operator
[fx] No. Please try again.
[f*] Correct!
[l]promoter
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]regulatory gene
[fx] No. Please try again.
[f*] Great!
[l]regulatory mRNA
[fx] No. Please try again.
[f*] Correct!
[l]regulatory protein
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]RNA polymerase
[fx] No. Please try again.
[f*] Correct!
[l]structural genes
[fx] No. Please try again.
[f*] Excellent!
[q labels = “top”]
[l]operator
[fx] No. Please try again.
[f*] Good!
[l]promoter
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]regulatory gene
[fx] No. Please try again.
[f*] Excellent!
[l]regulatory mRNA
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]regulatory protein
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]RNA polymerase
[fx] No. Please try again.
[f*] Excellent!
[l]structural genes
[fx] No, that’s not correct. Please try again.
[f*] Great!
[l]lactose
[fx] No. Please try again.
[f*] Correct!
[l]lactose-digesting enzymes
[fx] No, that’s not correct. Please try again.
[f*] Great!
[l]mRNA for lactose-digesting enzymes
[fx] No. Please try again.
[f*] Excellent!
[x][restart]
[/qwiz]
5. The Trp Operon
An organism’s metabolic activities can be divided into categories: breaking things down (often to release energy, as in cellular respiration) and building things up. The Lac operon is in the first category: it produces enzymes for the breakdown of the disaccharide lactose into the two monosaccharides; glucose and galactose.
The Trp operon is in the second category: it’s a system for controlling the synthesis of tryptophan, one of the 20 amino acids that make up proteins.
Because the Trp operon involves synthesis, the logic behind its control is the opposite of that involved in the Lac operon. For the trp operon, the logic is as follows: if tryptophan is lacking in the environment, then the operon system should be turned on so that the cell can synthesize the tryptophan it needs. On the other hand, if tryptophan is available, then the operon should be turned off: instead of wasting energy and matter making tryptophan, the cell should simply let tryptophan diffuse in, and use it for whatever protein it’s synthesizing.
We’ll start with what happens when tryptophan is in E. coli’s environment (and thus diffusing into the cell). And based on what you know about the lac operon, you should be able to figure out why, in this situation, the structural genes are not transcribed.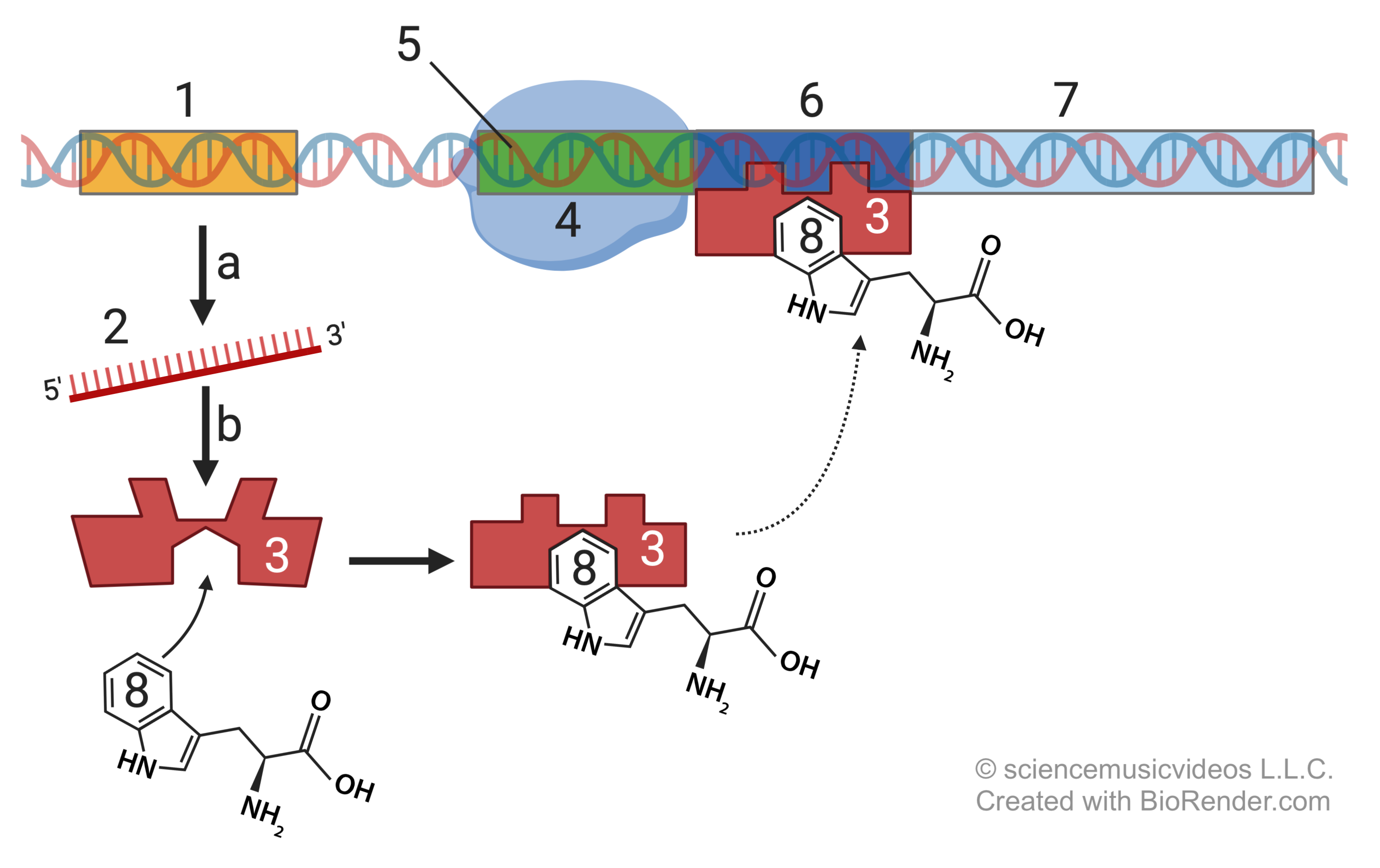
Here’s what’s happening. The regulatory gene (1) produces regulatory mRNA (2), which gets translated into a regulatory protein (3).
This regulatory protein has two binding sites. If tryptophan is present in the environment, it will bind with the regulatory protein, which causes a change in the regulatory protein’s DNA binding site. In this case, the conformational change is such that the regulatory protein can bind with the operator (6). With the operator occupied, RNA polymerase (4) can’t move past the promoter (5), and can’t transcribe the structural genes (7).
In the trp operon, the regulatory protein is often referred to as a repressor. Because the regulatory protein only binds to the operator when tryptophan is present, tryptophan is often referred to as a co-repressor.
If tryptophan is not present in the environment, the situation changes as shown below.
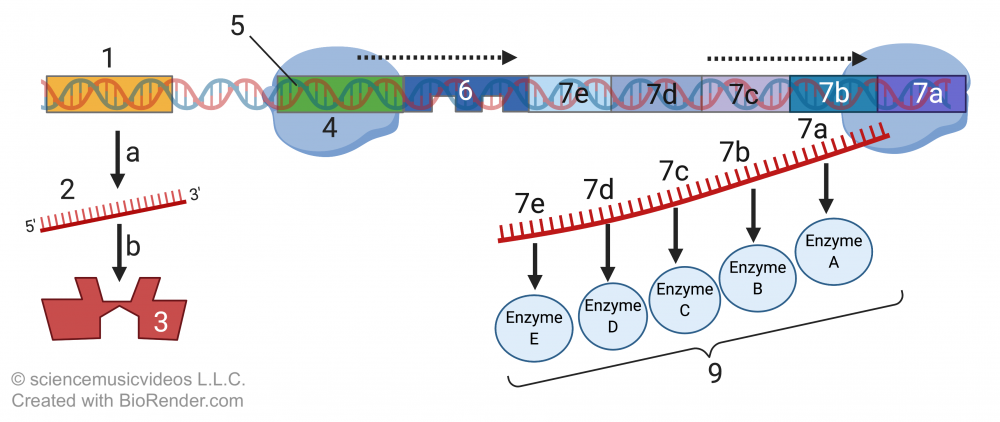
Without tryptophan, the regulatory protein (3) can’t bind with the operator (6). RNA polymerase (4) is free to bind with the operon’s promoter (5) and transcribe the structural genes (7A through 7E), producing the structural mRNA (8A through 8E) that will be translated into the structural proteins (9A through 9E) that constitute the pathway for synthesizing tryptophan.
Got it? Label the diagrams below.
6. Trp operon labeled diagrams
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Tryp Operon Labeled Diagrams (M16) “]
[h]Trp operon: Labeled Diagrams
[i]
[q labels = “top”]
[l]enzymes for tryptophan synthesis
[fx] No, that’s not correct. Please try again.
[f*] Correct!
[l]mRNA for structural genes
[fx] No. Please try again.
[f*] Good!
[l]operator
[fx] No. Please try again.
[f*] Good!
[l]promoter
[fx] No. Please try again.
[f*] Great!
[l]regulatory gene
[fx] No. Please try again.
[f*] Good!
[l]regulatory protein
[fx] No. Please try again.
[f*] Good!
[l]mRNA for regulatory protein
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]RNA polymerase
[fx] No. Please try again.
[f*] Good!
[l]structural genes
[fx] No. Please try again.
[f*] Good!
[q labels = “top”]
[l]operator
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]promoter
[fx] No, that’s not correct. Please try again.
[f*] Correct!
[l]regulatory gene
[fx] No. Please try again.
[f*] Correct!
[l]regulatory gene’s mRNA
[fx] No. Please try again.
[f*] Great!
[l]regulatory protein
[fx] No. Please try again.
[f*] Good!
[l]RNA polymerase
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]structural genes
[fx] No. Please try again.
[f*] Excellent!
[l]tryptophan
[fx] No. Please try again.
[f*] Good!
[/qwiz]
7. Operons Quiz
This quiz uses diagrams and multiple-choice questions to test your understanding of operons. Good luck!
[qwiz qrecord_id=”sciencemusicvideosMeister1961-Operons Quiz (M16)”]
[h]Operons
[i]Biohaiku
An operator
Blocked by repressor protein
Prevents transcription
[q json=”true” dataset_id=”Operons|1d4690ab006f00″ question_number=”1″] Which number is the regulatory gene?
[textentry single_char=”true”]
[c]ID E=[Qq]
[f]IFllcy4g4oCcMeKAnSBpcyB0aGUgcmVndWxhdG9yecKgZ2VuZS4=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBUaGUgcmVndWxhdG9yeSBnZW5lIGNvZGVzIGZvciBhIHJlZ3VsYXRvcnkgcHJvdGVpbiwgd2hpY2ggY2FuLCBpbiB0dXJuLCBiaW5kIHdpdGggYSByZWdpb24gb24gdGhlIEROQSBjYWxsZWQgYW4gb3BlcmF0b3IuIFNvLCBmaW5kIHNvbWV0aGluZyB0aGF0IHJlcHJlc2VudHMgc29tZXRoaW5nIHRoYXQgY291bGQgYmluZCB3aXRoIHRoZSBETkEsIHRoZW4gdXNlIHRoZSBhcnJvd3MgaW4gdGhlIGRpYWdyYW0gdG8gZmluZCB0aGUgRE5BIHRoYXQgY29kZXMgZm9yIHRoYXQgaXQu
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d463f2d604300″ question_number=”2″] Which number is the regulatory mRNA?
[textentry single_char=”true”]
[c]ID I=[Qq]
[f]IFllcy4g4oCcMuKAnSBpcyB0aGUgcmVndWxhdG9yeSBtUk5BLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBSZW1lbWJlciB0aGUgY2VudHJhbCBkb2dtYSB0aGF0IEROQSBtYWtlcyBSTkEgbWFrZXMgcHJvdGVpbi4gWW91IGNhbiBzZWUgdGhpcyBwcm9jZXNzIHJlcHJlc2VudGVkIG9uIHRoZSBsZWZ0IHNpZGUgb2YgdGhlIGRpYWdyYW0uIEluIHRlcm1zIG9mIHRoaXMgcXVlc3Rpb24sIHJlbWVtYmVyIHRoYXQgdGhlIHJlZ3VsYXRvcnkgbVJOQSBjb2RlcyBmb3IgYSByZWd1bGF0b3J5IHByb3RlaW4sIHdoaWNoIGNhbiwgaW4gdHVybiwgYmluZCB3aXRoIGEgcmVnaW9uIG9uIHRoZSBETkEgY2FsbGVkIGFuIG9wZXJhdG9yLiBTbywgZmluZCBzb21ldGhpbmcgdGhhdCByZXByZXNlbnRzIHNvbWV0aGluZyB0aGF0IGNvdWxkIGJpbmQgd2l0aCB0aGUgRE5BLCB0aGVuIHVzZSB0aGUgYXJyb3dzIGluIHRoZSBkaWFncmFtIHRvIGZpbmQgdGhlIG1STkEgdGhhdCBjb2RlcyBmb3IgaXQu
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d45eb5bb43300″ question_number=”3″] Which number is the regulatory protein?
[textentry single_char=”true”]
[c]ID M=[Qq]
[f]IFllcy4g4oCcM+KAnSBpcyB0aGUgcmVndWxhdG9yeSBwcm90ZWluLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBSZW1lbWJlciB0aGUgY2VudHJhbCBkb2dtYSB0aGF0IEROQSBtYWtlcyBSTkEgbWFrZXMgcHJvdGVpbi4gWW91IGNhbiBzZWUgdGhpcyBwcm9jZXNzIHJlcHJlc2VudGVkIG9uIHRoZSBsZWZ0IHNpZGUgb2YgdGhlIGRpYWdyYW0uIFRoZSBwcm90ZWluIHdvdWxkIGJlIGF0IHRoZSBlbmQgb2YgdGhpcyBjaGFpbiBvZiBldmVudHMuIEluIGFkZGl0aW9uLCB0aGlzIHByb3RlaW4gY2FuIGJpbmQgd2l0aCB0aGUgRE5BIGF0IGEgcmVnaW9uIGNhbGxlZCB0aGUgb3BlcmF0b3IuIFVzZSB0aGVzZSBjbHVlcyB0byBmaW5kIHRoZSBhbnN3ZXIgbmV4dCB0aW1lLg==
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d45a32e439700″ question_number=”4″] Which number is RNA polymerase?
[textentry single_char=”true”]
[c]ID Q=[Qq]
[f]IFllcy4g4oCcNOKAnSBpcyBSTkEgcG9seW1lcmFzZS4=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBSTkEgcG9seW1lcmFzZSBpcyBhIHByb3RlaW4gdGhhdCB0cmFuc2NyaWJlcyBETkEgaW50byBtUk5BLiBJZiAmIzgyMjA7N2EmIzgyMjE7IHRocm91Z2ggJiM4MjIwOzdjJiM4MjIxOyByZXByZXNlbnQgdGhpcyBvcGVyb24mIzgyMTc7cyBzdHJ1Y3R1cmFsIGdlbmVzLCBhbmQgOSByZXByZXNlbnRzIG1STkEsIHdoYXQmIzgyMTc7cyB0aGUgb25seSB0aGluZyBpbiB0aGlzIGRpYWdyYW0gdGhhdCBjb3VsZCBiZSB0cmFuc2NyaWJpbmcgdGhlIGZvcm1lciBpbnRvIHRoZSBsYXR0ZXI/
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d455d54dedf00″ question_number=”5″] Which number is the promoter?
[textentry single_char=”true”]
[c]ID U=[Qq]
[f]IFllcy4g4oCcNeKAnSBpc8KgdGhlIHByb21vdGVyLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LsKgVGhlIHByb21vdGVyIGlzIHRoZSByZWdpb24gd2hlcmUgUk5BIHBvbHltZXJhc2UgYmluZHMgdG8gYmVnaW4gdHJhbnNjcmlwdGlvbiBvZiB0aGUgb3Blcm9uJiM4MjE3O3Mgc3RydWN0dXJhbCBnZW5lcy4gSWYgUk5BIHBvbHltZXJhc2UgaXMgJiM4MjIwOzQsJiM4MjIxOyB0aGVuIHdoYXQgaGFzIHRvIGJlIHRoZSBwcm9tb3Rlcj8=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d4512d3625f00″ question_number=”6″] Which number is the operator?
[textentry single_char=”true”]
[c]ID Y=[Qq]
[f]IFllcy4g4oCcNuKAnSBpc8KgdGhlIG9wZXJhdG9yLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBUaGUgb3BlcmF0b3IgaXMganVzdCAmIzgyMjA7ZG93bnN0cmVhbSYjODIyMTsgb2YgdGhlIHByb21vdGVyIGFuZCBqdXN0ICYjODIyMDt1cHN0cmVhbSYjODIyMTsgb2YgdGhlIHN0cnVjdHVyYWwgZ2VuZXMuIElmIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzIGFyZSBhdCA3YSB0aHJvdWdoIDdjLCBhbmQgdGhlIHByb21vdGVyIGlzIGF0ICYjODIyMDs0LCYjODIyMTsgdGhlbiB3aGljaCBudW1iZXIgaXMgcmVwcmVzZW50aW5nIHRoZSBvcGVyYXRvcj8=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d44c851e5df00″ question_number=”7″ multiple_choice=”true”] Which of the following would represent one of the structural genes?
[c]IDE=[Qq]
[f]IE5vLiAmIzgyMjA7MSYjODIyMTsgcmVwcmVzZW50cyBhIHJlZ3VsYXRvcnkgZ2VuZS4=[Qq]
[c]IDU=[Qq]
[f]IE5vLiAmIzgyMjA7NSYjODIyMTsgcmVwcmVzZW50cyB0aGUgcHJvbW90ZXIu[Qq]
[c]IDY=[Qq]
[f]IE5vLiAmIzgyMjA7NiYjODIyMTsgcmVwcmVzZW50cyB0aGUgb3BlcmF0b3Iu[Qq]
[c]ID di[Qq]
[f]IFllcy4gJiM4MjIwOzdiJiM4MjIxOyBpcyBvbmUgb2YgdGhyZWUgc3RydWN0dXJhbCBnZW5lcyBpbiB0aGUgbGFjIG9wZXJvbi4gRWFjaCBvZiB0aGVzZSBnZW5lcyBnZXRzIHRyYW5zY3JpYmVkIGludG8gbVJOQSwgYW5kIHRoZW4gc3ludGhlc2l6ZWQgaW50byBvbmUgb2YgdGhlIHRocmVlIHByb3RlaW5zIHRoYXQgYnJlYWsgZG93biBsYWN0b3NlIGludG8gZ2x1Y29zZSBhbmQgZ2FsYWN0b3NlLg==
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d447dd0695f00″ question_number=”8″] Which number represents lactose?
[textentry single_char=”true”]
[c]ID g=[Qq]
[f]IFllcy4g4oCcOOKAnSBpcyBsYWN0b3NlLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlIGFyZSB0d28gaGludHMuIDEpIExhY3Rvc2UgaXMgYSBkaXNhY2NoYXJpZGUsIGEgc3VnYXIgbWFkZSBvZiB0d28gc2ltcGxlIHN1Z2FyIChtb25vc2FjY2hhcmlkZSkgc3VidW5pdHMuIFdoYXQgb24gdGhpcyBkaWFncmFtIGNvdWxkIGJlIG1hZGUgb2YgdHdvIHN1YnVuaXRzPyDCoDIpIExhY3Rvc2UgYmluZHMgd2l0aCB0aGUgcmVndWxhdG9yeSBwcm90ZWluLCB3aGljaCB0dXJucyB0aGUgb3Blcm9uICYjODIyMDtvZmYuJiM4MjIxOyBJZiAmIzgyMjA7MyYjODIyMTsgaXMgdGhlIHJlZ3VsYXRvcnkgcHJvdGVpbiwgdGhlbiB3aGF0IGNvdWxkIGJlIHNvbWV0aGluZyB0aGF0IGJpbmRzIHdpdGggdGhpcyBwcm90ZWluPw==
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d4435a2f8c300″ question_number=”9″] Which number represents the mRNA that’s transcribed from the structural genes in this operon?
[textentry single_char=”true”]
[c]ID k=[Qq]
[f]IFllcy4g4oCcOeKAnSBpcyB0aGUgbVJOQSB0aGF0IGNvZGVzIGZvciB0aGUgb3Blcm9uJiM4MjE3O3Mgc3RydWN0dXJhbCBnZW5lcy4=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBJZiB0aGUgdGhyZWUgYm94ZXMgYXQgMTAgcmVwcmVzZW50ZWQgdGhlIGdlbmUgcHJvZHVjdHMsIGFuZCB0aGUgdGhyZWUgZ2VuZXMgYXQgJiM4MjIwOzcmIzgyMjE7IHJlcHJlc2VudCB0aGUgc3RydWN0dXJhbCBnZW5lcywgdGhlbiB3aGF0IGhhcyB0byBiZSB0aGUgbVJOQSBmb3IgdGhlc2UgZ2VuZXM/
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d43eb217c4300″ question_number=”10″ multiple_choice=”true”] Which of the following would best describe the operon shown below?
[c]IGluZHVjaWJsZc KgIMKgIMKgIA==[Qq][c]IHJlcHJlc3NpYmxl
Cg==[Qq][f]IFllcy4gVGhlIG9wZXJvbiBzaG93biBhYm92ZSBpcyAmIzgyMjA7aW5kdWNpYmxlLiYjODIyMTsgVG8gJiM4MjIwO2luZHVjZSYjODIyMTsgaXMgdG8gY2F1c2Ugc29tZXRoaW5nIHRvIGhhcHBlbi4gVGhlIGxhYyBvcGVyb24gaXMgaW5kdWNpYmxlIGJlY2F1c2UgbGFjdG9zZSwgd2hlbiBwcmVzZW50LCB0dXJucyB0aGUgb3Blcm9uICYjODIyMDtvbi4mIzgyMjE7IFRoZSBkZWZhdWx0IGNvbmRpdGlvbiwgd2hlbiBsYWN0b3NlIGlzIGFic2VudCwgaXMgZm9yIHRoaXMgb3Blcm9uIHN5c3RlbSB0byBiZSB0dXJuZWQgJiM4MjIwO29mZi4mIzgyMjE7[Qq]
[f]IE5vLiBUaGUgb3Blcm9uIHNob3duIGFib3ZlIGlzICYjODIyMDtpbmR1Y2libGUuJiM4MjIxOyBUbyAmIzgyMjA7aW5kdWNlJiM4MjIxOyBpcyB0byBjYXVzZSBzb21ldGhpbmcgdG8gaGFwcGVuLiBUaGUgbGFjIG9wZXJvbiBpcyBpbmR1Y2libGUgYmVjYXVzZSBsYWN0b3NlLCB3aGVuIHByZXNlbnQsIHR1cm5zIHRoZSBvcGVyb24gJiM4MjIwO29uLiYjODIyMTsgVGhlIGRlZmF1bHQgY29uZGl0aW9uLCB3aGVuIGxhY3Rvc2UgaXMgYWJzZW50LCBpcyBmb3IgdGhpcyBvcGVyb24gc3lzdGVtIHRvIGJlIHR1cm5lZCAmIzgyMjA7b2ZmLiYjODIyMTs=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d439e4bf3df00″ question_number=”11″ multiple_choice=”true”] The diagram below represents the lac operon. Which statement about the diagram below is most correct?
[c]IFRoZSBzdHJ1Y3R1cmFsIGdlbmVzIGFyZSBiZWluZyB0cmFuc2NyaWJlZCBpbnRvIGEgcHJvdGVpbiwgYXMgaXMgc2hvd24gYXQgJiM4MjIwOzEsJiM4MjIxOyAmIzgyMjA7MiwmIzgyMjE7IGFuZCAmIzgyMjA7My4mIzgyMjE7[Qq]
[f]IE5vLiBXaGF0JiM4MjE3O3Mgc2hvd24gYXQgJiM4MjIwOzEsJiM4MjIxOyAmIzgyMjA7MiwmIzgyMjE7IGFuZCAmIzgyMjA7MyYjODIyMTsgYXJlIHRoZSByZWd1bGF0b3J5IGdlbmUsIHRoZSBtUk5BIHRyYW5zY3JpYmVkIGZyb20gdGhpcyBnZW5lLCBhbmQgdGhlIHJlZ3VsYXRvcnkgcHJvdGVpbi4gVGhlIHN0cnVjdHVyYWwgZ2VuZXMgKGF0ICYjODIyMDs3JiM4MjIxOykgYXJlIGJsb2NrZWQgYmVjYXVzZSB0aGUgcmVndWxhdG9yeSBwcm90ZWluICgzKSBpcyBib3VuZCB0byB0aGUgb3BlcmF0b3IgKDYpLg==[Qq]
[c]IFRoZSBzdHJ1Y3R1cmFsIGdlbmVzIGFyZSBub3QgYmVpbmcgZXhwcmVzc2Vk IGJlY2F1c2UgbGFjdG9zZSBpcyBub3QgaW4gdGhlIGVudmlyb25tZW50Lg==[Qq]
[f]IFllcy4gQmVjYXVzZSBsYWN0b3NlIGlzIG5vdCBpbiB0aGUgZW52aXJvbm1lbnQsIHRoZSByZWd1bGF0b3J5IHByb3RlaW4gKDMpIGhhcyBhIGNvbmZvcm1hdGlvbiB0aGF0IGVuYWJsZXMgaXQgdG8gYmluZCB3aXRoIHRoZSBvcGVyYXRvciAoNiksIHByZXZlbnRpbmcgUk5BIHBvbHltZXJhc2UgZnJvbSB0cmFuc2NyaWJpbmcgdGhlIHN0cnVjdHVyYWwgZ2VuZXMu[Qq]
[c]IFRoZSBzdHJ1Y3R1cmFsIGdlbmVzIGFyZSBub3QgYmVpbmcgZXhwcmVzc2VkLCBiZWNhdXNlIFJOQSBwb2x5bWVyYXNlIGlzIGJsb2NraW5nIHRoZSByZWd1bGF0b3J5IHByb3RlaW4gZnJvbSB0cmFuc2NyaWJpbmcgdGhlIGdlbmVzLg==[Qq]
[f]IE5vIChidXQgeW91IGhhdmUgdGhlIHJpZ2h0IGlkZWEpLiBCZWNhdXNlIGxhY3Rvc2UgaXMgbm90IGluIHRoZSBlbnZpcm9ubWVudCwgdGhlIHJlZ3VsYXRvcnkgcHJvdGVpbiAoMykgaGFzIGEgY29uZm9ybWF0aW9uIHRoYXQgZW5hYmxlcyBpdCB0byBiaW5kIHdpdGggdGhlIG9wZXJhdG9yICg2KSwgcHJldmVudGluZyBSTkEgcG9seW1lcmFzZSBmcm9tIHRyYW5zY3JpYmluZyB0aGUgc3RydWN0dXJhbCBnZW5lcy4=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d42769a0dc300″ question_number=”15″] Which number represents the substrate of the enzymes produced by this operon?
[textentry single_char=”true”]
[c]ID g=[Qq]
[f]IFllcy4g4oCcOOKAnSBpcyBsYWN0b3NlLCB3aGljaCBpcyB0aGUgc3Vic3RyYXRlIG9mIHRoZSBlbnp5bWVzIHRoYXQgdGhpcyBvcGVyb24gcHJvZHVjZXMu[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYcKgaGludC7CoFRoZSBzdWJzdHJhdGUgb2YgdGhlIGVuenltZXMgcHJvZHVjZWQgYnkgdGhpcyBvcGVyb24gaXMgdGhlIHNhbWUgbW9sZWN1bGUgd2hvc2UgcHJlc2VuY2UgdHVybnMgdGhlIG9wZXJvbiAmIzgyMjA7b24uJiM4MjIxOyBXaGljaCBtb2xlY3VsZSBiaW5kcyB3aXRoIHRoZSByZWd1bGF0b3J5IHByb3RlaW4gKHNob3duIGF0ICYjODIyMDszJiM4MjQyOykgaW4gYSB3YXkgdGhhdCBjaGFuZ2VzIHRoYXQgcHJvdGVpbiYjODIxNztzIHNoYXBlLCBrZWVwaW5nIGl0IGZyb20gYmluZGluZyB3aXRoIHRoZSBvcGVyYXRvcj8=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d41e63f2c8b00″ question_number=”17″] The diagram below represents the trp operon. Which number is the regulatory mRNA?
[textentry single_char=”true”]
[c]ID I=[Qq]
[f]IFllcy4g4oCcMuKAnSBpcyB0aGUgcmVndWxhdG9yeSBtUk5BLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBSZW1lbWJlciB0aGUgY2VudHJhbCBkb2dtYSB0aGF0IEROQSBtYWtlcyBSTkEgbWFrZXMgcHJvdGVpbi4gWW91IGNhbiBzZWUgdGhpcyBwcm9jZXNzIHJlcHJlc2VudGVkIG9uIHRoZSBsZWZ0IHNpZGUgb2YgdGhlIGRpYWdyYW0uIEluIHRlcm1zIG9mIHRoaXMgcXVlc3Rpb24sIHJlbWVtYmVyIHRoYXQgdGhlIHJlZ3VsYXRvcnkgbVJOQSBjb2RlcyBmb3IgYSByZWd1bGF0b3J5IHByb3RlaW4sIHdoaWNoIGNhbiwgaW4gdHVybiwgYmluZCB3aXRoIGEgcmVnaW9uIG9uIHRoZSBETkEgY2FsbGVkIGFuIG9wZXJhdG9yLiBTbywgZmluZCBzb21ldGhpbmcgdGhhdCByZXByZXNlbnRzIHNvbWV0aGluZyB0aGF0IGNvdWxkIGJpbmQgd2l0aCB0aGUgRE5BLCB0aGVuIHVzZSB0aGUgYXJyb3dzIGluIHRoZSBkaWFncmFtIHRvIGZpbmQgdGhlIG1STkEgdGhhdCBjb2RlcyBmb3IgaXQu[Qq]
[q json=”true” dataset_id=”Operons|1d40b094ff1700″ question_number=”21″] The diagram below represents the trp operon. Which number is the operator?
[textentry single_char=”true”]
[c]ID Y=[Qq]
[f]IFllcy4g4oCcNuKAnSBpc8KgdGhlIG9wZXJhdG9yLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50LiBUaGUgb3BlcmF0b3IgaXMganVzdCAmIzgyMjA7ZG93bnN0cmVhbSYjODIyMTsgb2YgdGhlIHByb21vdGVyIGFuZCBqdXN0ICYjODIyMDt1cHN0cmVhbSYjODIyMTsgb2YgdGhlIHN0cnVjdHVyYWwgZ2VuZXMuIElmIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzIGFyZSBhdCA3LCBhbmQgdGhlIHByb21vdGVyIGlzIGF0ICYjODIyMDs0LCYjODIyMTsgdGhlbiB3aGljaCBudW1iZXIgaXMgcmVwcmVzZW50aW5nIHRoZSBvcGVyYXRvcj8=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d4016e9ee4f00″ question_number=”23″] The diagram below represents the trp operon. Which number represents tryptophan?
[textentry single_char=”true”]
[c]ID g=[Qq]
[f]IFllcy4g4oCcOOKAnSBpcyB0cnlwdG9waGFuLg==[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50OiB0cnlwdG9waGFuIGlzIHdoYXQmIzgyMTc7cyBzZWVuIGJpbmRpbmcgdG8gdGhlIHJlZ3VsYXRvcnkgcHJvdGVpbiAoJiM4MjIwOzMmIzgyNDM7KSwgY2hhbmdpbmcgaXRzIHNoYXBlIHNvIHRoYXQgaXQgY2FuIGJpbmQgdG8gdGhlIG9wZXJhdG9yIHJlZ2lvbiBvZiB0aGUgb3Blcm9uLg==
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3f7aead1a300″ question_number=”25″] Shown below is a representation of the tryp operon from the Biology Coloring Workbook. Which letter represents the operator region of the operon?
[textentry single_char=”true”]
[c]IE Q=[Qq]
[f]IFllcy4g4oCcRCYjODIyMTvCoGlzIHRoZSBvcGVyYXRvci4=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBGaW5kIGEgbGV0dGVyIHRoYXQgcmVwcmVzZW50cyB0aGUgcGFydCBvZiB0aGUgb3Blcm9uJiM4MjE3O3MgRE5BIHRoYXQgdGhlIHJlcHJlc3NvciBwcm90ZWluIChKKSBjYW4gYmluZCB0bywgYmxvY2tpbmcgdHJhbnNjcmlwdGlvbi4=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3edeebb4f700″ question_number=”27″] Which number represents this operon’s co-repressor?
[textentry single_char=”true”]
[c]ID g=[Qq]
[f]IFllcy4g4oCcOOKAnSBpcyB0aGUgY28tcmVwcmVzc29yLCB0cnlwdG9waGFuLiBJdCYjODIxNztzIGNhbGxlZCBhIGNvLXJlcHJlc3NvciBiZWNhdXNlIGl0IHdvcmtzIHdpdGggdGhlIHJlcHJlc3NvciAoJiM4MjIwOzMmIzgyMjE7KSB0byBiaW5kIHdpdGggdGhlIG9wZXJhdG9yIHRvIHNodXQgZG93biB0aGUgdHJhbnNjcmlwdGlvbiBvZiB0aGUgc3RydWN0dXJhbCBnZW5lcy4=[Qq]
[c]ICo=[Qq]
[f]IE5vLiBIZXJlJiM4MjE3O3MgYSBoaW50OiB0aGUgY28tcmVwcmVzc29ywqBpcyB3aGF0JiM4MjE3O3Mgc2VlbiBiaW5kaW5nIHRvIHRoZSByZWd1bGF0b3J5IHByb3RlaW4gKCYjODIyMDszJiM4MjQzOyksIGNoYW5naW5nIGl0cyBzaGFwZSBzbyB0aGF0IGl0IGNhbiBiaW5kIHRvIHRoZSBvcGVyYXRvciByZWdpb24gb2YgdGhlIG9wZXJvbi4gSW4gb3RoZXIgd29yZHMsIHlvdSYjODIxNztyZSBsb29raW5nIGZvciB0cnlwdG9waGFuLg==
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3e8d6e14cb00″ question_number=”28″ multiple_choice=”true”] In the lac operon system
[c]IFJlcHJlc3NvciBwcm90ZWlucyBtYXkgYmluZCB0byB0aGUgb3BlcmF0b3IgcmVnaW9u LCBibG9ja2luZyB0cmFuc2NyaXB0aW9uIG9mIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzLg==[Qq]
[f]IENvcnJlY3QuIEluIHRoZSBsYWMgb3Blcm9uLCB3aGVuIHJlcHJlc3NvciBwcm90ZWlucyBhcmUgYm91bmQgdG8gdGhlIG9wZXJhdG9yLCB0aGUgc3RydWN0dXJhbCBnZW5lcyBjYW5ub3QgYmUgdHJhbnNjcmliZWQu[Qq]
[c]IHN0cnVjdHVyYWwgZ2VuZXMgbWFrZSBhIHJlcHJlc3NvciBwcm90ZWluIHRoYXQgYmluZCB0byB0aGUgcHJvbW90ZXIsIHByZXZlbnRpbmcgdHJhbnNjcmlwdGlvbi4=[Qq]
[f]IE5vLiBUaGUgc3RydWN0dXJhbCBnZW5lcyBtYWtlIHRoZSBlbnp5bWVzIGZvciBkaWdlc3RpbmcgbGFjdG9zZS4=[Qq]
[c]IGxhY3Rvc2UgdHVybnMgdGhlIG9wZXJvbiBvZmY=[Qq]
[f]IE5vLiBJbiB0aGUgbGFjIG9wZXJvbiwgbGFjdG9zZSB0dXJucyB0aGUgb3Blcm9uICYjODIyMDtvbi4mIzgyMjE7IEluIG90aGVyIHdvcmRzLCB0aGUgcnVsZSBpcyAmIzgyMjA7b25seSBtYWtlIHRoZSBlbnp5bWVzIGZvciBsYWN0b3NlIGRpZ2VzdGlvbiB3aGVuIGxhY3Rvc2UgaXMgaW4gdGhlIGVudmlyb25tZW50LiYjODIyMTs=[Qq]
[c]IHJpYm9zb21lcyBjYW5ub3QgcGFzcyBieSB0aGUgcmVwcmVzc29yIHByb3RlaW4gYm91bmQgdG8gdGhlIG9wZXJhdG9yIHJlZ2lvbg==[Qq]
[f]IE5vLiBJdCYjODIxNztzIFJOQSBwb2x5bWVyYXNlIChub3Qgcmlib3NvbWVzKSB0aGF0IGNhbiYjODIxNzt0IHBhc3MgYnkgdGhlIHJlcHJlc3NvciBwcm90ZWluIGJvdW5kIHRvIHRoZSBvcGVyYXRvciByZWdpb24u
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3e2950157f00″ question_number=”29″ multiple_choice=”true”] In an operon, active repressor molecules have an effect when they bind to:
[c]IFByb21vdGVycw==[Qq]
[f]IE5vLiBSTkEgcG9seW1lcmFzZSBpcyB3aGF0IGJpbmRzIHRvIHRoZSBwcm9tb3Rlci4gVGhpcyBxdWVzdGlvbiBpcyBhc2tpbmcgeW91IHdoYXQgcGFydCBvZiB0aGUgb3Blcm9uIHN5c3RlbSByZXByZXNzb3IgbW9sZWN1bGVzIGJpbmQgdG8uIExvb2sgY2xvc2VseSBhdCB0aGUgd29yZCAmIzgyMjA7b3Blcm9uJiM4MjIxOyBmb3IgYSBjbHVlLg==[Qq]
[c]IFJpYm9zb21lcw==[Qq]
[f]IE5vLiBSaWJvc29tZXMgZG9uJiM4MjE3O3QgcGxheSBhbnkgcm9sZSBhcyByZXByZXNzb3JzIHdpdGhpbiBvcGVyb25zICh0aG91Z2ggdGhleSBkbyB0cmFuc2xhdGUgdGhlIHJlZ3VsYXRvcnkgYW5kIHN0cnVjdHVyYWwgUk5BKS7CoFRoaXMgcXVlc3Rpb24gaXMgYXNraW5nIHlvdSB3aGF0IHBhcnQgb2YgdGhlIG9wZXJvbiBzeXN0ZW0gcmVwcmVzc29yIG1vbGVjdWxlcyBiaW5kIHRvLiBMb29rIGNsb3NlbHkgYXQgdGhlIHdvcmQgJiM4MjIwO29wZXJvbiYjODIyMTsgZm9yIGEgY2x1ZS4=[Qq]
[c]IFN0cnVjdHVyYWwgZ2VuZXM=[Qq]
[f]IE5vLiBSZXByZXNzb3IgbW9sZWN1bGVzIGJsb2NrIHRyYW5zY3JpcHRpb24gb2YgdGhlIHN0cnVjdHVyYWwgZ2VuZXMsIGJ1dCBub3QgYnkgZGlyZWN0bHkgYmluZGluZyB0byB0aGVtLsKgTG9vayBjbG9zZWx5IGF0IHRoZSB3b3JkICYjODIyMDtvcGVyb24mIzgyMjE7IGZvciBhIGNsdWUu[Qq]
[c]IFJOQSBwb2x5bWVyYXNl[Qq]
[f]IE5vLiBUaGUgcmVwcmVzc29yIG1vbGVjdWxlcyBibG9jayBSTkEgcG9seW1lcmFzZSwgYnV0IHRoZXkgZG9uJiM4MjE3O3QgYmluZCB0byBpdC7CoFRoaXMgcXVlc3Rpb24gaXMgYXNraW5nIHlvdSB3aGF0IHBhcnQgb2YgdGhlIG9wZXJvbiBzeXN0ZW0gcmVwcmVzc29yIG1vbGVjdWxlcyBiaW5kIHRvLiBMb29rIGNsb3NlbHkgYXQgdGhlIHdvcmQgJiM4MjIwO29wZXJvbiYjODIyMTsgZm9yIGEgY2x1ZS4=[Qq]
[c]IE9wZXJh dG9ycw==[Qq]
[f]IFllcyEgSW4gYW4gb3Blcm9uLCByZXByZXNzb3IgbW9sZWN1bGVzIGhhdmUgYW4gZWZmZWN0IHdoZW4gdGhleSBiaW5kIHRvIG9wZXJhdG9ycy4=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3dd0d651a700″ question_number=”30″ multiple_choice=”true”] The lac operon system is an important adaptation for E. coli bacteria because the sugar lactose:
[c]IElzIHJlcXVpcmVkIGZvciB0aGUgcHJvZHVjdGlvbiBvZiB0aGUgc3RydWN0dXJhbCBnZW5lcyB0aGF0IGNyZWF0ZSB0aGUgcmVwcmVzc29yIHByb3RlaW5zIHVwb24gd2hpY2ggdGhpcyBvcGVyb24gZGVwZW5kcy4=[Qq]
[f]IE5vLiBGaXJzdCwgbGFjdG9zZSBpcyB0aGUgc3Vic3RyYXRlIG9mIHRoZSBlbnp5bWVzIHByb2R1Y2VkIGJ5IHRoZSBsYWMgb3Blcm9uIHN5c3RlbTogaXQmIzgyMTc7cyBub3QgcHJvZHVjZWQgYnkgdGhlIHN5c3RlbS4gU2Vjb25kLCBpdCYjODIxNztzIG5vdCBwYXJ0IG9mIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzICh3aGljaCBhcmUgbWFkZSBvZiBETkEpLg==[Qq]
[c]IElzIHRoZSBvbmx5IHN1Z2FyIHRoYXQgRS4gY29sacKgY2FuIHVzZQ==[Qq]
[f]IE5vLiBFLiBjb2xpwqBjYW4gbWV0YWJvbGl6ZSBvdGhlciBzdWdhcnMgYXMgd2VsbCwgbm90YWJseSBnbHVjb3NlLg==[Qq]
[c]IElzIG9ubHnCoGF2YWlsYWJsZSBzb21ldGltZXM7IG1ha2luZyBsYWN0b3NlLWRpZ2VzdG luZyBlbnp5bWVzIGNvbnRpbnVvdXNseSB3b3VsZCBiZSBhIHdhc3RlIG9mIGVuZXJneQ==[Qq]
[f]IFllcy4gTGFjdG9zZSBpcyBvbmx5IHBlcmlvZGljYWxseSBhdmFpbGFibGUgdG8gRS4gY29saSwgc28gaXQgb25seSBtYWtlcyBzZW5zZSBmb3IgdGhlc2UgYmFjdGVyaWEgdG8gc3ludGhlc2l6ZSB0aGUgZW56eW1lcyBmb3IgZGlnZXN0aW5nIGxhY3Rvc2Ugd2hlbiBpdCYjODIxNztzIGF2YWlsYWJsZSBpbiB0aGUgZW52aXJvbm1lbnQu[Qq]
[c]IEFjdHMgdG8gdHVybiBvZmYgdGhlIHN5c3RlbQ==[Qq]
[f]IE5vLiBJbiB0aGUgTGFjIG9wZXJvbiBzeXN0ZW0sIHRoZSBwcmVzZW5jZSBvZiBsYWN0b3NlIHR1cm5zIHRoZSBzeXN0ZW0gb24uIEluIHRoZSBwcmVzZW5jZSBvZiBsYWN0b3NlLCB0aGUgc3RydWN0dXJhbCBnZW5lcyB0aGF0IGNvZGUgZm9yIGxhY3Rvc2UtZGlnZXN0aW5nIGVuenltZXMgYXJlIHRyYW5zY3JpYmVkIGFuZCB0cmFuc2xhdGVkIHNvIHRoYXQgRS4gY29saSBjYW4gZGlnZXN0IHRoaXMgZW5lcmd5IHNvdXJjZS4=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3d7d04a59700″ question_number=”31″ multiple_choice=”true”] The structural genes in the lac operon system are only transcribed when
[c]IExhY3Rvc2UgaXMgYWJzZW50[Qq]
[f]IE5vLiBJbiB0aGUgbGFjIG9wZXJvbiwgdGhlIHN0cnVjdHVyYWwgZ2VuZXMgYXJlIHRyYW5zY3JpYmVkIHdoZW4gbGFjdG9zZSBpcyBwcmVzZW50Lg==[Qq]
[c]IFRoZSByZXByZXNzb3IgcHJvdGVpbiBpcyBhdHRhY2hlZCB0byB0aGUgb3BlcmF0b3IgZ2VuZQ==[Qq]
[f]IE5vLiBXaGVuIHRoZSByZXByZXNzb3IgcHJvdGVpbiBpcyBhdHRhY2hlZCB0byB0aGUgb3BlcmF0b3IgZ2VuZSwgdGhlIFJOQSBwb2x5bWVyYXNlIGlzIGJsb2NrZWQgYW5kIGNhbiYjODIxNzt0IHRyYW5zY3JpYmUgdGhlIHN0cnVjdHVyYWwgZ2VuZXMu[Qq]
[c]IExhY3Rvc2UgYmluZHMgdG8gdGhlIFJOQSBwb2x5bWVyYXNl[Qq]
[f]IE5vLiBUaGUgbGFjIG9wZXJvbiBzeXN0ZW0gd29ya3MgdGhyb3VnaCB0aGUgcmVwcmVzc29yIHByb3RlaW4gYmluZGluZyB0byB0aGUgb3BlcmF0b3IgcmVnaW9uLCB3aGljaCBibG9ja3MgUk5BIHBvbHltZXJhc2UgZnJvbSB0cmFuc2NyaWJpbmcgdGhlIHN0cnVjdHVyYWwgZ2VuZXMuIE5vdGhpbmcgaW4gdGhpcyBzeXN0ZW0gYmluZHMgZGlyZWN0bHkgdG8gUk5BIHBvbHltZXJhc2Uu[Qq]
[c]IExhY3Rvc2UgYmluZHMgd2l0aCB0 aGUgcmVwcmVzc29yIHByb3RlaW4=[Qq]
[f]IEV4Y2VsbGVudC4gSW4gdGhlIGxhYyBvcGVyb24gc3lzdGVtLCB0aGUgc3RydWN0dXJhbCBnZW5lcyBhcmUgb25seSB0cmFuc2NyaWJlZCB3aGVuIGxhY3Rvc2UgYmluZHMgdG8gdGhlIHJlcHJlc3NvciBwcm90ZWluLiBUaGlzIGNhdXNlcyBhbiBhbGxvc3RlcmljIGNoYW5nZSBpbiB0aGUgc2hhcGUgb2YgdGhlIHJlcHJlc3NvciBwcm90ZWluIHNvIHRoYXQgaXQgY2FuIG5vIGxvbmdlciBiaW5kIHRvIHRoZSBvcGVyYXRvciByZWdpb24sIGFsbG93aW5nIHRyYW5zY3JpcHRpb24gb2YgdGhlIHN0cnVjdHVyYWwgZ2VuZXMgdG8gb2NjdXIu[Qq]
[c]IEJhY3RlcmlhIG1ha2UgbGFjdG9zZQ==[Qq]
[f]IE5vLiBUaGUgbGFjIG9wZXJvbiBpcyBhYm91dCBob3cgYmFjdGVyaWEgZGlnZXN0IGxhY3Rvc2UuIE1hbW1hbHMgKGxpa2UgcGVvcGxlLCBjb3dzLCB3aGFsZXMsIGFuZCBiYXRzKSBtYWtlIGxhY3Rvc2Ugd2hlbiB0aGV5IG1ha2UgbWlsay4=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3d2932f98700″ question_number=”32″ multiple_choice=”true”] In the lac operon system in E. coli, the repressor molecules are:
[c]IExhY3Rvc2Ugc3VnYXJz[Qq]
[f]IE5vLiBMYWN0b3NlIHN1Z2FycyBhcmUgdGhlIHN1YnN0cmF0ZSBvZiB0aGUgZW56eW1lcyBwcm9kdWNlZCBieSB0aGUgc3RydWN0dXJhbCBnZW5lcy4gV2hhdCBraW5kIG9mIG1vbGVjdWxlIGNvdWxkIGJsb2NrIHRoZSB0cmFuc2NyaXB0aW9uIG9mIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzPw==[Qq]
[c]IFByb3 RlaW5z[Qq]
[f]IFllcyEgVGhlIHJlcHJlc3NvciBtb2xlY3VsZXMgYXJlIHByb3RlaW5zIHByb2R1Y2VkIGJ5IHRyYW5zY3JpcHRpb24gYW5kIHRyYW5zbGF0aW9uIG9mIHRoZSBvcGVyb24mIzgyMTc7cyByZWd1bGF0b3J5IGdlbmVzLg==[Qq]
[c]IFNob3J0IHBpZWNlcyBvZiBETkE=[Qq]
[f]IE5vLiBUaGUgcmVwcmVzc29yIG1vbGVjdWxlcyBpbiB0aGXCoGxhYyBvcGVyb24gc3lzdGVtIGFyZSBjYXBhYmxlIG9mIGJpbmRpbmcgdG8gRE5BIGFuZCBibG9ja2luZyB0cmFuc2NyaXB0aW9uIG9mIHRoZSBvcGVyb24mIzgyMTc7cyBzdHJ1Y3R1cmFsIGdlbmVzIChidXQgdGhleSYjODIxNztyZSBub3QgbWFkZSBvZiBETkEpLiBXaGF0wqBraW5kIG9mIHJlZ3VsYXRvcnkgbW9sZWN1bGUgY291bGQgYmluZCB0byBETkEgaW4gdGhpcyB3YXk/[Qq]
[c]IEJvdW5kIHRvIHRoZSBwcm9tb3RvciByZWdpb24=[Qq]
[f]IE5vLCBUaGUgcmVwcmVzc29yIG1vbGVjdWxlcyBpbiB0aGXCoGxhYyBvcGVyb24gc3lzdGVtIGJpbmQgdG8gdGhlIG9wZXJhdG9yIHJlZ2lvbiwgd2hpY2ggaXMganVzdCBkb3duc3RyZWFtIG9mIHRoZSBwcm9tb3Rlci4gV2hhdCBraW5kIG9mIG1vbGVjdWxlcyBjb3VsZCBhIGNlbGwgcHJvZHVjZSB0aGF0IGNvdWxkIGJpbmQgdG8gRE5BIGluIHRoaXMgd2F5Pw==[Qq]
[c]IFByb2R1Y2VkIGJ5IGEgc3RydWN0dXJhbCBnZW5l[Qq]
[f]IE5vLsKgVGhlIHJlcHJlc3NvciBtb2xlY3VsZXMgaW4gdGhlIGxhYyBvcGVyb24gc3lzdGVtIGFyZSBwcm9kdWNlZCBieSB0cmFuc2NyaXB0aW9uIGFuZCB0cmFuc2xhdGlvbiBvZiB0aGUgcmVndWxhdG9yeSBnZW5lLiBXaGF0IGtpbmQgb2YgbW9sZWN1bGVzIGFyZSB0aGUgcHJvZHVjdCBvZiB0cmFuc2NyaXB0aW9uIGFuZCB0cmFuc2xhdGlvbj8=
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3cce6529cb00″ question_number=”33″ multiple_choice=”true”] In the lac operon system, the presence of lactose causes E. coli cells to produce a series of enzymes that break lactose apart into two monosaccharides that can then be further metabolized for matter and energy. When E. coli runs out of lactose:
[c]IHRoZSByZXByZXNzb3IgcHJvdGVpbiBiaW5kcyB0byB0aGUgcHJvbW90ZXIgcmVnaW9u[Qq]
[f]IE5vLiBJbiB0aGUgbGFjIG9wZXJvbiBzeXN0ZW0sIHdoZW4gRS4gY29saSBydW5zIG91dCBvZiBsYWN0b3NlLCB0aGUgcmVwcmVzc29yIHByb3RlaW4gYXNzdW1lcyBhIHNoYXBlIHRoYXQgZW5hYmxlcyBpdCB0byBiaW5kIHdpdGggdGhlIG9wZXJhdG9yIHJlZ2lvbiAobm90IHRoZSBwcm9tb3Rlciku[Qq]
[c]IHRoZSByZXByZXNzb3IgcHJvdGVpbiBiaW 5kcyB0byB0aGUgb3BlcmF0b3IgcmVnaW9u[Qq]
[f]IFllcy7CoEluIHRoZSBsYWMgb3Blcm9uIHN5c3RlbSwgd2hlbiBFLiBjb2xpIHJ1bnMgb3V0IG9mIGxhY3Rvc2UsIHRoZSByZXByZXNzb3IgcHJvdGVpbiBhc3N1bWVzIGEgc2hhcGUgdGhhdCBlbmFibGVzIGl0IHRvIGJpbmQgd2l0aCB0aGUgb3BlcmF0b3IgcmVnaW9uLCBwcmV2ZW50aW5nIFJOQSBwb2x5bWVyYXNlIGZyb20gdHJhbnNjcmliaW5nIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzLg==[Qq]
[c]IFJOQSBwb2x5bWVyYXNlIGF0dGFjaGVzIHRvIHRoZSByZXByZXNzb3IgcmVnaW9u[Qq]
[f]IE5vLiBPcGVyb25zIGhhdmUgb3BlcmF0b3IgcmVnaW9ucywgbm90IHJlcHJlc3NvciByZWdpb25zLiDCoEluIHRoZSBsYWMgb3Blcm9uIHN5c3RlbSwgd2hlbiBFLiBjb2xpIHJ1bnMgb3V0IG9mIGxhY3Rvc2UsIHRoZSByZXByZXNzb3IgcHJvdGVpbiBhc3N1bWVzIGEgc2hhcGUgdGhhdCBlbmFibGVzIGl0IHRvIGJpbmQgd2l0aCB0aGUgb3BlcmF0b3IsIHdoaWNoIGJsb2NrcyBSTkEgcG9seW1lcmFzZSBmcm9tIHRyYW5zY3JpYmluZyB0aGUgc3RydWN0dXJhbCBnZW5lcy4=[Qq]
[c]IFJOQSBwb2x5bWVyYXNlIGFsbG93cyB0aGUgc3RydWN0dXJhbCBnZW5lcyB0byBtYWtlIG1STkE=[Qq]
[f]IE5vLiBJdCYjODIxNztzIHRoZSBvcHBvc2l0ZS4gSW4gdGhlIGxhYyBvcGVyb24gc3lzdGVtLCB3aGVuIEUuIGNvbGkgcnVucyBvdXQgb2YgbGFjdG9zZSwgdGhlIHJlcHJlc3NvciBwcm90ZWluIGFzc3VtZXMgYSBzaGFwZSB0aGF0IGVuYWJsZXMgaXQgdG8gYmluZCB3aXRoIHRoZSBvcGVyYXRvciwgd2hpY2ggYmxvY2tzIFJOQSBwb2x5bWVyYXNlIGZyb20gdHJhbnNjcmliaW5nIHRoZSBzdHJ1Y3R1cmFsIGdlbmVzLg==
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3c7a937dbb00″ question_number=”34″ multiple_choice=”true”] The tryp operon system is an important adaptation for E. coli bacteria because the amino acid tryptophan
[c]IGlzIHBhcnRpY3VsYXJseSBpbXBvcnRhbnQgZm9yIA==RS4gY29saSYjODIxNztzwqBtZXRhYm9saXNtIChtdWNoIG1vcmUgc28sIGZvciBleGFtcGxlLCB0aGFuIHRoZSBhbWlubyBhY2lkcyBhcmdpbmluZSBvciBhc3BhcmFnaW5lKS4=[Qq]
[f]IE5vLsKgRS4gY29saQ==IGNlcnRhaW5seSBuZWVkcyB0aGUgYW1pbm8gYWNpZCB0cnlwdG9waGFuIHRvIHN5bnRoZXNpemUgcHJvdGVpbnMsIGJ1dCBubyBtb3JlIHRoYW4gYW55IG90aGVyIGFtaW5vIGFjaWQu[Qq]
[c]IGlzIHNvbWV0aW1lcyBhdmFpbGFibGUgaW4gdGhlIGVudmlyb25tZW50OyBtYWtpbmcg dHJ5cHRvcGhhbsKgY29udGludW91c2x5IHdvdWxkIGJlIGEgd2FzdGUgb2YgZW5lcmd5[Qq]
[f]IFllcy7CoEJlY2F1c2UgdHJ5cHRvcGhhbiBpcyBvY2Nhc2lvbmFsbHkgYXZhaWxhYmxlIGZyb20gdGhlIGVudmlyb25tZW50LMKgbWFraW5nIHRyeXB0b3BoYW7CoGNvbnRpbnVvdXNseSB3b3VsZCBiZSBhIHdhc3RlIG9mIGVuZXJneQ==[Qq]
[c]IGFjdHMgdG8gdHVybiBvbiB0aGUgdHJhbnNjcmlwdGlvbiBvZiB0aGUgc3RydWN0dXJhbCBnZW5lcyBmb3IgdHJ5cHRvcGhhbiBzeW50aGVzaXMu[Qq]
[f]IE5vLsKgSW4gdGhlIHRyeXAgb3Blcm9uIHN5c3RlbSwgdHJ5cHRvcGhhbiYjODIxNztzIHByZXNlbmNlIHR1cm5zIG9mZiB0aGUgdHJhbnNjcmlwdGlvbiBvZiB0aGUgc3RydWN0dXJhbCBnZW5lcy4=[Qq]
[c]IGlzIGFuIGltcG9ydGFudCBlbmVyZ3kgc291cmNlIGZvciBFLiBjb2xp[Qq]
[f]IE5vLiBZb3UgbWlnaHQgYmUgdGhpbmtpbmcgYWJvdXQgdGhlIGxhYyBvcGVyb24sIHdoaWNoIGlzIGFib3V0IGRpZ2VzdGluZyB0aGUgc3VnYXIgbGFjdG9zZSwgYSBzb3VyY2Ugb2YgZW5lcmd5IGFuZCBtYXR0ZXIgZm9yIEUuIGNvbGkuIFRoZSB0cnlwwqBvcGVyb24gc3lzdGVtIGlzIGFib3V0IEUuIGNvbGkmIzgyMTc7cyBhYmlsaXR5IHRvIGNvbnRyb2wgbWV0YWJvbGljIHBhdGh3YXlzIGZvciBzeW50aGVzaXppbmcgbmVlZGVkIHN1YnN0YW5jZXMgKGluIHRoaXMgY2FzZSwgdGhlIGFtaW5vIGFjaWQgdHJ5cHRvcGhhbiku
Cg==[Qq]
[q json=”true” dataset_id=”Operons|1d3c1fc5adff00″ question_number=”35″ multiple_choice=”true”] The structural genes in the trp operon system are only transcribed when
[c]IHRyeXB0b3BoYW4gaXMgcHJlc2VudCBhbmQgYmluZHMgd2l0aCB0aGUgcmVwcmVzc29yIHByb3RlaW4=[Qq]
[f]IE5vLiBJbiB0aGUgdHJ5cMKgb3Blcm9uLCB0aGUgcmVndWxhdG9yeSBzY2hlbWUgaXMgYWJvdXQgYXZvaWRpbmcgc3ludGhlc2l6aW5nIGEgbmVlZGVkIHN1YnN0YW5jZSB3aGVuIHRoYXQgc3Vic3RhbmNlIGlzIGZyZWVseSBhdmFpbGFibGUgZnJvbSB0aGUgZW52aXJvbm1lbnQuIEluIG90aGVyIHdvcmRzLCB0aGUgc3RydWN0dXJhbCBnZW5lcyAod2hpY2ggY29kZSBmb3IgZW56eW1lcyB0aGF0IHN5bnRoZXNpemUgdHJ5cHRvcGhhbikgYXJlIG9ubHkgdHJhbnNjcmliZWQgd2hlbiB0cnlwdG9waGFuwqBpcyBhYnNlbnQu[Qq]
[c]IFRoZSByZXByZXNzb3IgcHJvdGVpbiBpcyBhdHRhY2hlZCB0byB0aGUgb3BlcmF0b3IgZ2VuZQ==[Qq]
[f]IE5vLiBXaGVuIHRoZSByZXByZXNzb3IgcHJvdGVpbiBpcyBhdHRhY2hlZCB0byB0aGUgb3BlcmF0b3IgZ2VuZSwgdGhlIFJOQSBwb2x5bWVyYXNlIGlzIGJsb2NrZWQgYW5kIGNhbiYjODIxNzt0IHRyYW5zY3JpYmUgdGhlIHN0cnVjdHVyYWwgZ2VuZXMuIEluIHRoZSB0cnlwIG9wZXJvbiwgdGhpcyBvbmx5IG9jY3VycyBpbiB0aGUgYWJzZW5jZSBvZiB0cnlwdG9waGFuLg==[Qq]
[c]IHRyeXB0b3BoYW4gYmluZHMgdG8gUk5BIHBvbHltZXJhc2U=[Qq]
[f]IE5vLiBUaGUgdHJ5cMKgb3Blcm9uIHN5c3RlbSB3b3JrcyB0aHJvdWdoIHRoZSByZXByZXNzb3IgcHJvdGVpbiBiaW5kaW5nIHRvIHRoZSBvcGVyYXRvciByZWdpb24sIHdoaWNoIGJsb2NrcyBSTkEgcG9seW1lcmFzZSBmcm9tIHRyYW5zY3JpYmluZyB0aGUgc3RydWN0dXJhbCBnZW5lcy4gTm90aGluZyBpbiB0aGlzIHN5c3RlbSBiaW5kcyBkaXJlY3RseSB0byBSTkEgcG9seW1lcmFzZS4=[Qq]
[c]IHRoZSBjby1yZXByZXNzb3IgKHRy eXB0b3BoYW4pIGlzIGFic2VudC4=[Qq]
[f]IEV4Y2VsbGVudC4gSW4gdGhlIHRyeXDCoG9wZXJvbiBzeXN0ZW0sIHRyeXB0b3BoYW4gYWN0cyBhcyBhIGNvLXJlcHJlc3Nvci4gSXQmIzgyMTc7cyBvbmx5IHdoZW4gdHJ5cHRvcGhhbiBpcyBhYnNlbnQgdGhhdCB0aGUgc3RydWN0dXJhbCBnZW5lcyBpbiB0aGlzIG9wZXJvbiAod2hpY2ggY29kZSBmb3IgZW56eW1lcyB0aGF0IHN5bnRoZXNpemUgdHJ5cHRvcGhhbikgYXJlIHRyYW5zY3JpYmVkLg==[Qq]
[/qwiz]
8. ADVANCED TOPIC: CAP (Catabolite Activator Protein) and Positive Gene Regulation
For the AP Biology exam, the material on operons shown above is probably sufficient. But there’s another level to understanding prokaryotic gene regulation: If you’re interested, go for it!
We’ve seen above how the Lac operon allows E. coli to turn on the production of enzymes to digest lactose when lactose is in its environment and to turn the production of these same enzymes off when lactose is not in the environment. But there’s an even more subtle regulatory scheme. If an E. coli cell is simultaneously fed with both lactose and glucose, it will preferentially use glucose, and only turn to lactose when glucose is in short supply. From an energetic perspective, this makes sense. Glucose is the simplest carbohydrate: the genes for metabolizing it are always turned “on.” Lactose, on the other hand, requires special enzymes to process. Given the choice, it’s energetically favorable for the cell to break down glucose first. But what system allows E. coli to express this preference?
The preference for glucose is controlled through the expression of another molecule: cyclic AMP (cAMP). We’ve met cAMP before, as a key second messenger in cell communication. In the context of operons, here’s how cAMP works.
Let’s consider a situation where both glucose (shown below by the letter “J”) and lactose (shown below at “G”) are present in the cell’s environment.
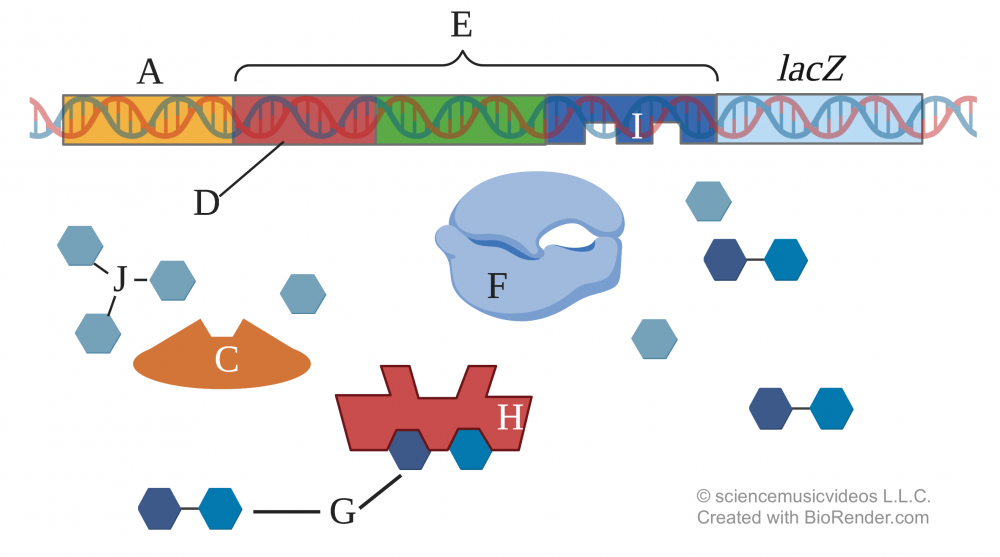
Under these circumstances, another regulatory protein called Catabolite Activator Protein (CAP, shown at “C” above) has a shape that doesn’t allow it to bind with the Lac Promoter (“E”) at the CAP binding site (“D”). CAP enhances the ability of RNA polymerase to bind with the promoter. So even though lactose (“G”) is present, and even though the Lac regulatory protein (“H”) won’t bind with the Lac operator (“I”), very little transcription of the structural genes will occur. Glucose, not lactose, is the fuel that will be metabolized.
But if the glucose level drops and the lactose level stays high, then everything changes.
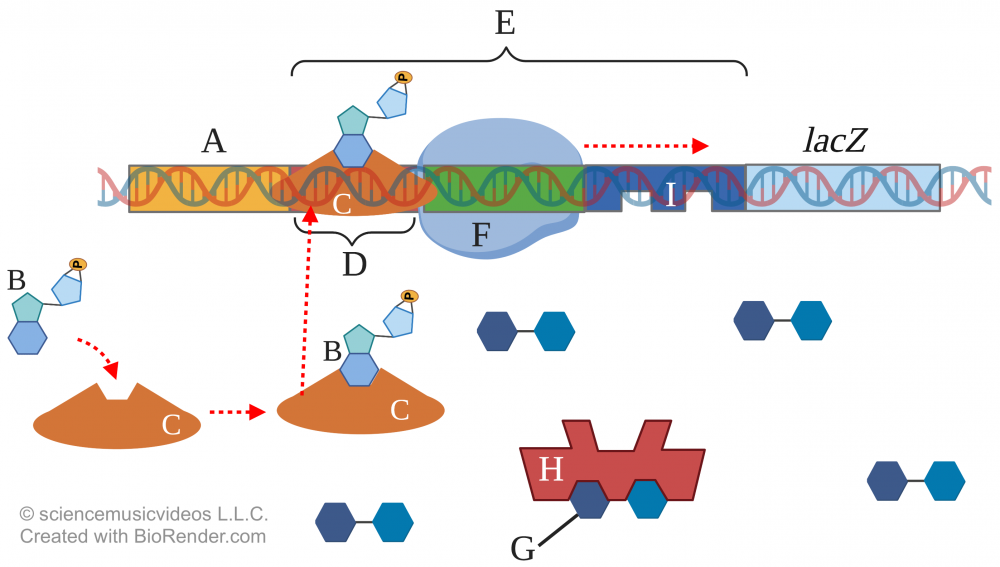
The low level of glucose induces the production of cAMP (“B”). cAMP binds with CAP (at “C”) changing its conformation in such a way that it 1) binds with the promoter region of the lac operon at the CAP binding site (“D”). This binding enhances RNA polymerase’s ability to bind with the promoter (“E”). And with the regulatory protein (“H”) allosterically altered so that it won’t bind with the operator (“I”) RNA polymerase can transcribe the structural genes (7) for lactose digesting enzymes (not shown above).
As the authors of Campbell Biology conclude
Thus the lac operon is under dual control: negative control by the lac repressor and positive control by CAP. The state of the lac repressor…determines whether or not transcription occurs…; the state of CAP controls the rate of transcription…(Campbell Biology, 9th edition, page 355).
Got it? Label the diagrams below.
9. cAMP/CAP labeled diagrams
[qwiz qrecord_id=”sciencemusicvideosMeister1961-CAP (Positive Gene Regulation Labeled Diagrams (M16)”]
[h]Positive Gene Regulation in Prokaryotes: cAMP and CAP
[i]
[q labels = “top”]
[l]CAP binding site
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]Catabolite Activator Protein
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]glucose
[fx] No. Please try again.
[f*] Excellent!
[l]lactose
[fx] No. Please try again.
[f*] Correct!
[l]operator
[fx] No. Please try again.
[f*] Correct!
[l]Promoter
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]regulatory gene
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]regulatory protein
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]RNA polymerase
[fx] No, that’s not correct. Please try again.
[f*] Great!
[l]structural genes
[fx] No. Please try again.
[f*] Correct!
[q labels = “top”]
[l]CAP binding site
[fx] No. Please try again.
[f*] Good!
[l]Catabolite Activator Protein (inactive)
[fx] No, that’s not correct. Please try again.
[f*] Good!
[l]Catabolite Activator Protein (active)
[fx] No, that’s not correct. Please try again.
[f*] Great!
[l]lactose
[fx] No, that’s not correct. Please try again.
[f*] Excellent!
[l]operator
[fx] No. Please try again.
[f*] Excellent!
[l]Promoter
[fx] No. Please try again.
[f*] Good!
[l]regulatory gene
[fx] No. Please try again.
[f*] Correct!
[l]regulatory protein
[fx] No, that’s not correct. Please try again.
[f*] Great!
[l]RNA polymerase
[fx] No, that’s not correct. Please try again.
[f*] Great!
[l]structural genes
[fx] No. Please try again.
[f*] Correct!
[l]cAMP
[fx] No. Please try again.
[f*] Great!
[/qwiz]
What’s next?
This tutorial ends this module on Bacterial Genetics and Operons. Use the menus above to choose another tutorial.

